LEARNING AND MEMORY PATHWAY INHIBITORS T-5224 AND KN-62 ON THE EXPRESSION OF C-FOS AND TASTE RECEPTOR T1R1 IN SINIPERCA CHUATSI
-
摘要: 为探究翘嘴鳜 (Siniperca chuatsi) 中原癌基因c-Fos (proto-oncogene c-Fos)对摄食相关的味觉受体t1r1(taste receptor type 1 member 1, t1r1)的调控作用, 研究采用学习记忆通路相关抑制剂T-5224、KN-62处理鳜脑细胞, 通过不同浓度抑制剂处理发现T-5224对c-fos的起负调控作用而KN-62对c-fos起正调控作用, 同时筛选出适合的处理时间及浓度。30 μmol/L T-5224处理24h, 鳜脑中c-fos及t1r1的mRNA表达显著降低(P<0.05), 而200 nmol/L KN-62处理2h, 鳜脑中c-fos及t1r1的mRNA表达却极显著升高(P<0.05)。此外, 研究还分析了抑制剂处理翘嘴鳜脑细胞后, t1r1基因DNA甲基化变化。然而在两种抑制剂处理后, 鳜脑中t1r1的甲基化水平无显著变化(P>0.05)。以上结果说明, 翘嘴鳜脑细胞味觉受体t1r1转录水平变化, 可能是通过学习记忆因子c-fos基因调控, 而c-fos可能通过其他途径调控t1r1的转录, 但不是DNA甲基化。同时为翘嘴鳜驯化后摄食偏好变化提供理论依据。Abstract: Animals can improve their feeding efficiency and preference through learning and memory. As a key factor of learning and memory, c-fos plays an important role in animal feeding behavior. In this study, the brain cells of Siniperca chuatsi were treated with the inhibitors of learning and memory pathway T-5224 and KN-62. It was found that T-5224 had a negative regulation effect on c-fos while KN-62 had a positive regulation effect on c-fos, and the appropriate treatment time and concentration were selected. The mRNA expression of c-fos and t1r1 in the brain of mandarin fish decreased significantly after being treated with 30 μmol/L T-5224 for 24 hours (P<0.05), and increased significantly after being treated with 200 nmol/L KN-62 for 2h (P<0.01). In addition, the methylation changes of t1r1 gene after inhibitor treatment were also analyzed. However, there was no significant change in the methylation level of t1r1 in the brain of Siniperca chuatsi (P>0.05). The above results indicated that the transcription level of taste receptor t1r1 in brain cells of Siniperca chuatsi may be regulated by learning and memory factor c-fos gene, while c-fos may regulate t1r1 transcription through other ways, but not DNA methylation. At the same time, it provides theoretical basis for the change of feeding preference of Siniperca chuatsi after domestication. In this study, it was confirmed for the first time that T-5224 negatively regulated the learning and memory factor c-fos, while KN-62 positively regulated it. At the same time, the above results indicated that the expression of c-fos, the key transcription factor of learning and memory, affected the expression of taste receptor t1r1, and the c-fos gene might participate in the feeding habits of Siniperca chuatsi.
-
Keywords:
- Learning and memory pathway /
- Inhibitor /
- c-fos /
- t1r1 /
- DNA methylation /
- Siniperca chuatsi
-
学习和记忆是所有动物普遍存在的特征。动物通过学习和记忆使其灵活地适应不断变化的环境条件, 其中重要的是通过学习记忆获得新的觅食技能[1]和食物偏好[2-4]。动物的学习记忆过程涉及动物的多个行为过程, 包括摄食行为、集群行为、个体识别及信息传递等[5-7]。尽管学习记忆研究主要是在哺乳动物和鸟类中[8], 但也有研究报道, 鱼类中各种行为活动都受到学习记忆的影响[9]。在鱼类适应环境的过程中面临诸多挑战, 例如避免被捕食者捕食, 寻找食物及配偶选择等。为此鱼类凭借以往经验, 通过化学信号学习并判断新捕食者的威胁程度[10], 鱼类通过空间记忆和社会学习增强了对捕食者的规避能力。鱼类通过学习显著提高了摄食效率[3, 7], 还能增强鉴定合适配偶的能力[3, 11]。在这些鱼类行为中, 学习和记忆对鱼类的摄食行为中起着决定性的作用。最近的几项研究表明, 在大麻哈鱼(Oncorhynchus keta) [12]、孔雀鱼(Poecilia reticulata) [13]、褐鳟(Salmo trutta)[14]等鱼可以通过学习记忆提高摄食效率及摄食偏好, 而学习记忆能否在鱼类摄食偏好, 如驯食等过程中发挥作用, 目前尚不明确。
学习记忆相关基因之一的原癌基因c-Fos (proto-oncogene c-Fos) 的表达是小鼠(Mus musculus)海马区依赖性非空间记忆的形成和反复记忆所必需的[15]。有研究表明这c-fos基因的mRNA表达水平随着长期记忆等神经元激活反应而上升[16]。早期研究发现动物的T1R (taste receptor type 1, T1R) 基因家族缺失或突变, 以适应其特殊的摄食习性, 例如大熊猫(Ailuropoda melanoleuca)[17, 18]。以上研究表明动物能够通过位于味蕾上的鲜味受体感知食物的鲜味, 从而影响动物的摄食及食性偏好。有研究揭示团头鲂体内鲜味受体t1r1(taste receptor type 1 member 1, t1r1)缺失, 以适应植物食性[19]。通过草鱼转食前后的转录组数据发现, 鲜味受体t1r1的基因表达也发生了显著变化,
表明t1r1可能参与草鱼食性分化过程中的调控。然而, 学习记忆影响鱼类摄食及食性的过程中是否存在学习记忆相关基因c-fos对味觉受体t1r1表达的调控以及如何调控, 仍不清楚。有报道称外界环境的变化可以通过表观遗传调控, 尤其是DNA甲基化, 影响基因的转录, 最终改变鱼类的表型性状。例如通过基因控制区CpG岛的甲基化来调节基因转录的启动[20, 21]。
翘嘴鳜(Siniperca chuatsi)食性奇特,自鱼苗开口起就以活鱼虾为食, 经过长期的驯化后才可以接受人工饲料[22], 而长期驯食过程中的学习记忆促进了翘嘴鳜食性从活鱼转变为饲料[23]。何珊等[24]对杂交鳜驯食性状的转录组分析, 发现学习通路中c-fos基因差异表达。同时实验室前期研究发现易驯食鳜与易驯食饲料鳜脑中味觉受体t1r1的mRNA表达及其启动子甲基化模式与不易驯食鳜完全不同[25]。因此, 本实验以翘嘴鳜为研究对象, 利用学习记忆通路相关抑制剂T-5224和KN-62处理鳜脑细胞, 研究c-fos是否对味觉受体基因c-fos的表达存在调控作用及调控方式, 为进一步探讨学习记忆对鱼类摄食调控的分子机制, 并为改进翘嘴鳜人工饲料驯化技术提供理论依据。
1. 材料与方法
1.1 实验材料
实验动物: 实验用鱼为翘嘴鳜(体重100 g左右)取自华中农业大学鳜鱼研究中心, 用于后续分离翘嘴鳜脑细胞。实验开始之前先将翘嘴鳜转移到循环水养殖系统内暂养1周, 鱼缸水体体积为100 L, 曝气水养殖, 水体pH维持在6.8—7.2, 溶解氧含量维持在6.8—7.8 mg/L, 每日正常投喂饵料鱼, 换水吸污, 以维持翘嘴鳜稳定正常生理状态。
主要试剂: DL 2000 Marker(TaKaRa)、胎牛血清 (Fatal bovine serum, FBS)、胰蛋白酶、L15培养液 (Genom)、I型胶原酶 (Gibco)、两性霉素B、硫酸庆大霉素、青链霉素双抗 (HyClone 公司)、DPBS、Dimethyl sulfoxide (DMSO Sigma)、T-5224 (MedChemExpress)、KN-62 (MedChemExpress)
1.2 分离翘嘴鳜脑细胞
选取一尾状态比较好的翘嘴鳜, 用75%的酒精消毒, 在超净工作台里剖取脑组织于 DPBS 中清洗, 用剪刀镊子去除多余的组织, 然后将脑组织放入盛有AIM液的培养皿内浸泡消毒1.5h; 吸掉培养皿中多余AIM液, 用剪刀将脑组织剪成1 mm3大小的组织块, 将组织块转移到盛有胶原酶消化液的离心管中在28℃培养箱中消化2h; 加入L15培养液轻轻吹打得到脑细胞, 1000 r/min 室温离心5min后弃上清, 随后再用L15培养液漂洗两次(1000 r/min, 室温离心5min), 最后将收集的脑细胞加入原代培养液重悬后接种到细胞培养瓶, 于 28℃无CO2培养箱中培养, 第4天更换原代培养液1次。在显微镜下观察脑细胞的生长情况。本课题组已经采用Neu-N和β-tubulin免疫荧光细胞化学技术鉴定了脑细胞神经元纯度[26]。在脑原代细胞分离至第三代之后, 即进行实验。将脑细胞扩大培养, 铺板至6孔板中, 每个孔盛放2.0 mL培养液, 待细胞长满后进行下一步实验。
1.3 实验设计及样品采集
本实验设置对照组与实验组, 对照组加入抑制剂的溶剂, 实验组加入不同浓度抑制剂。抑制剂用DMSO稀释至适宜浓度, 当鳜脑原代细胞分离至第三代之后, 即可进行实验。将脑细胞扩大培养, 铺板至6 cm2平皿中, 每个平皿盛放2.5 mL培养液, 封口膜封口放入28℃无CO2恒温培养箱中培养。大概4d细胞覆盖率可达80%, 使用无添加试剂的L15基础培养基对原代细胞进行饥饿16—24h。抑制剂用DMSO稀释至适宜浓度备用, 吸除废液后再用无添加试剂的L15基础培养基进行清洗。吸取添加了抑制剂或缓冲液的L15基础培养基刺激细胞(根据不同抑制剂的效用不同, 添加的抑制剂量不同, 本研究T-5224采用0、10、30和60 μmol/L处理浓度; KN-62采用200 nmol/L), 封口膜封口, 放入28℃无CO2恒温培养箱中, 根据不同抑制剂效用, 决定抑制剂的刺激时间,本研究T-5224处理24h,KN-62处理2h。刺激时间到达后分别收集细胞中RNA和DNA进行后续实验操作, 样品冻存于–80℃超低温冰箱。
1.4 翘嘴鳜相关基因的 mRNA 表达水平分析
利用 TRIzol Reagent 试剂 (TaKaRa) 来提取翘嘴鳜脑原代细胞的总RNA, 再用 HiScript II Q RT SuperMix for qPCR 逆转录试剂盒(诺唯赞)进行反转录得到cDNA。采用实时荧光定量 RT-PCR 方法测定脑细胞中c-fos和t1r1基因的mRNA表达水平。反应体系 (20 μL) 如下: SYBR Green Supermix 10 μL, F 0.4 μL, R 0.4 μL, cDNA 模板 1 μL, ddH2O 8.2 μL。反应程序如下: 95℃, 5min; 95℃, 10s; 退火温度30s和延伸30s, 共39个循环。每经过一个循环, 收集一次荧光强度信号, 采用优化的2–∆∆Ct法, 以rpl13a基因的定量表达水平为内参进行相对表达量分析, 每个样品重复6次。本试验所用到的荧光定量引物均采用Primer 5.0软件进行设计, 并由生工合成, 引物序列见表 1。
表 1 实时荧光定量 PCR 的引物序列Table 1. Nucleotide sequences of the primers for real-time PCR基因
Gene name引物
Primer序列
Sepuences (5′—3′)退火温度
Annealing temperature (℃)产物大小
Product size (bp)rpl13a rpl13a-F TATCCCCCCACCCTATGACA 59 100 rpl13a-R ACGCCCAAGGAGAGCGAACT c-fos c-fos-F CGATGATGTTTACCGCTTTC 60 88 c-fos-R TAGTATCCCAGATTGTCCC t1r1 t1r1-F CGAGAGTGCCAAAGGAGGAG 58 278 t1r1-R TGGAGAGCAGATGGACAGC t1r1 BSP t1r1 F AGGGTTAATATAGATATAGATAACGATAGA 52 298 BSP t1r1 R CAACTAAATAATCAATTAAAAAATACAC 1.5 翘嘴鳜t1r1基因DNA甲基化水平分析和亚硫酸氢盐硫化PCR (BSP)
将收集的DNA样品采用TIANamp Genomic DNA Kit (Tiangen, Beijing, China) 试剂盒说明书的标准步骤提取基因组DNA。采用EZ DNA Methylation Kit (Zymo Research, Irvine, CA, USA)试剂盒说明书的标准步骤进行基因组DNA的亚硫酸氢盐修饰。原理如下: 基因组DNA中甲基化胞嘧啶碱基不转变, 同时, 未甲基化胞嘧啶脱氨基并转变成尿嘧啶, 经亚硫酸氢盐硫化PCR (Bisulphite Sequencing Polymerase Chain Reaction, BSP)扩增后为胸腺嘧啶, 从而区别甲基化/未甲基化的两类胞嘧啶。从实验室翘嘴鳜全基因组中获得t1r1的全序列[27], 基因5′侧翼区的序列, 第一外显子区的序列, ATG上游3500 bp的序列, 提交至在线预测网站Methprimer (http://www.urogene.org/cgibin/methprimer/methprimer.cgi) 进行预测以获得CpG岛 (CGI) 和候选CpG位点。鱼类CpG岛无明确定义, 选择默认参数并根据情况进行调整, 默认搜索参数如下: CpG岛片段大小 (Island size) > 100 bp, GC碱基含量(GC Percent) > 50.0%, CpG岛观察值/预测值 (Observed/Expected, Obs/Exp) > 0.6。BSP引物由在线软件MethPrimer 14.0推荐, 然后用Primer 5.0检测是否合格, 比较后采用特异性高引物用于BSP的扩增(表 1)。采用Taq plus DNA Polymerase (Vazyme Biotech, Nanjing, China) 在Biometra Thermo cyclers (Biometra, Göttingen, Germany) 仪器上进行PCR。PCR反应循环参数为: 94℃预变性5min; 然后如下程序进行40个循环, 即94℃变性30s, 52℃(根据不同引物的退火温度)退火30s, 72℃延伸30s; 之后72℃延伸7min, 最终12℃降温10s后结束。采用Gel Purification Kit (Sangon, Shanghai, China) 试剂盒纯化PCR产物, 然后克隆至载体pMD18-T clone vector (TaKaRa, Tokyo, Japan) 中。在每个实验组中, 先随机选取2个样品进行预扩增及分析, 存在DNA甲基化后检测全部6个样品。每个样品随机选取5个阳性克隆送至上海生工 (ABI3730 测序仪, Applied Biosystems) 进行测序, 每个驯食实验组总共收集成功测序的30个阳性克隆, 经测序得到DNA序列。采用在线网站QUMA (QUantification tool for Methylation Analysis) (http://quma.cdb.riken.jp/) 将测序序列与亚硫酸氢盐修饰前原序列进行比对分析, 根据比对结果确定候选CpG位点是否发生甲基化以及甲基化水平。
1.6 统计分析
采用软件SPSS 22.0 (IBM, Chicago, IL, USA) 的Shapiro-Wilk法标准化数据, 进行单因素方差分析。检查方差齐性之后, 通过Duncan’s multiple range test (MRT) 法检测均值间差异。实验组组间候选CpG位点甲基化水平的差异使用2×2卡方检验(χ2 test)分析, 实验组组间基因表达水平的差异使用独立样本T检验分析, P<0.05判定显著差异, P<0.01判定为极显著差异。所有数据表示为平均值±SEM (平均值的标准误差)。
2. 结果
2.1 T-5224刺激鳜脑细胞后学习记忆因子c-fos及味觉受体t1r1 mRNA表达水平变化
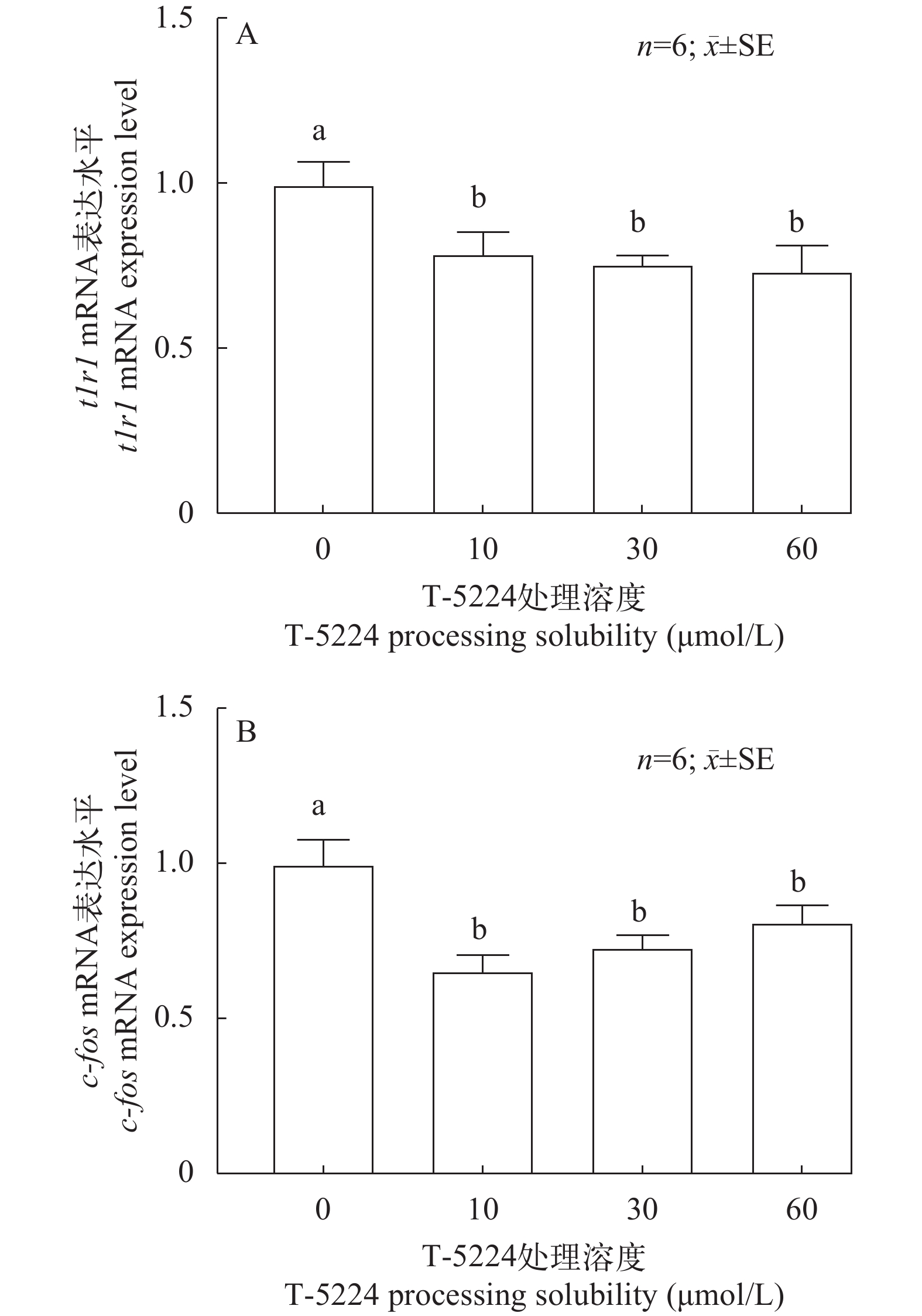
使用学习记忆因子c-fos的抑制剂T-5224处理鳜脑原代细胞, 利用RT-PCR技术比较使用T-5224抑制剂处理鳜脑细胞24h后, 对照组0和实验组n μmol/L c-fos和t1r1 (n表示不同浓度)mRNA水平(图 1)。不同的实验条件刺激鳜脑原代细胞得到不同的结果, 图 1A中不同浓度的T-5224处理与对照组相比, c-fos mRNA的表达水平均显著降低(P<0.05), 而10 μmol/L处理效果最明显, 30 μmol/L次之。与此同时在图 1B中不同浓度的T-5224处理与对照组相, t1r1 mRNA的表达水平均显著降低(P<0.05),而30和60 μmol/L处理效果最明显。
2.2 KN-62刺激鳜脑细胞后学习记忆因子c-fos及味觉受体t1r1 mRNA表达水平变化
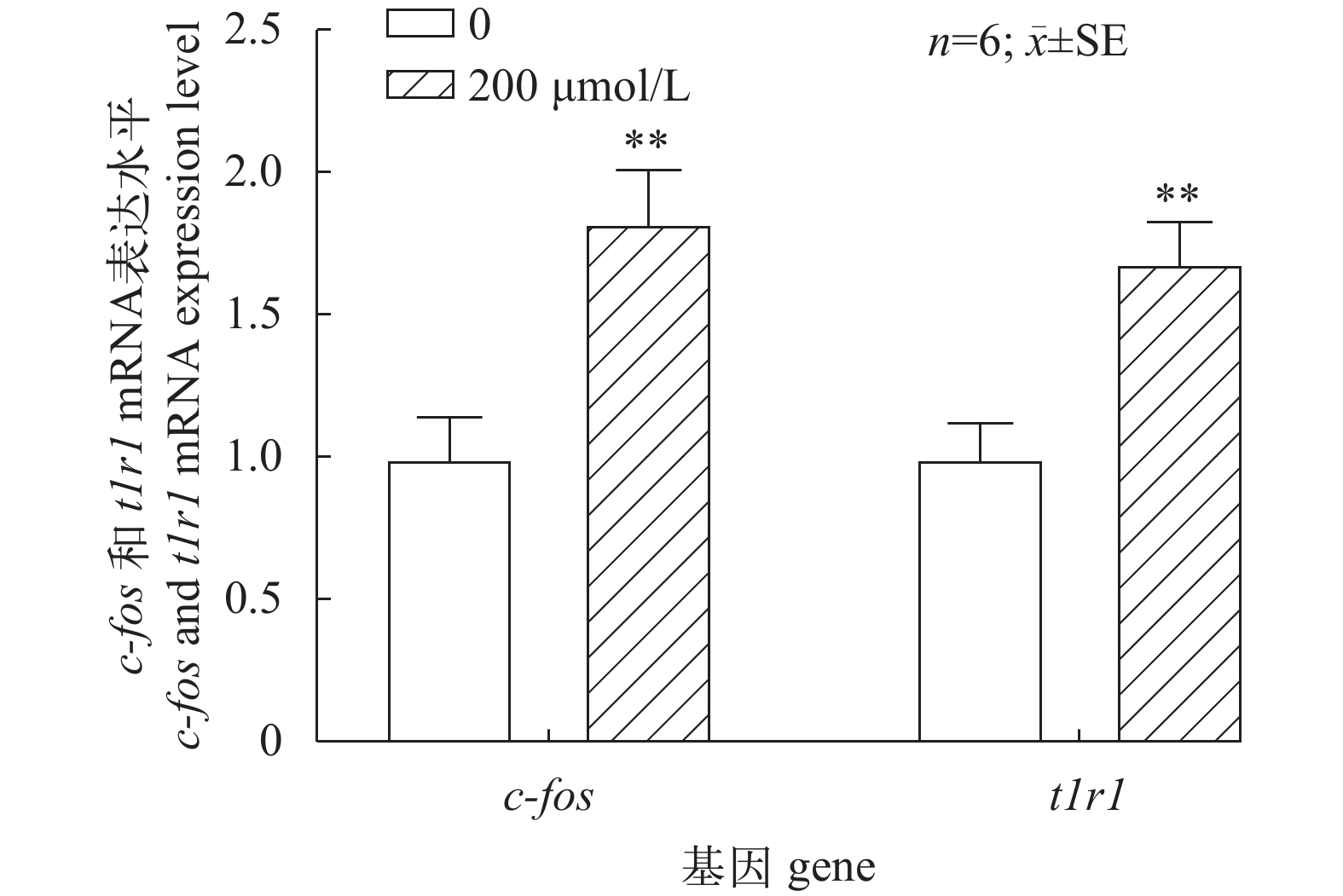
使用学习记忆通路CAMKII抑制剂KN-62处理鳜脑原代细胞, 参考彭建等[23]的选定最佳时间点和药物浓度, 我们选定2h这个时间进行对照组0和实验组200 nmol/L浓度抑制剂刺激实验(图 2), 在200 nmol/L-2h的刺激条件下, c-fos mRNA表达水平呈现极显著上升趋势 (P<0.01), 与此同时在相同处理条件下t1r1 mRNA表达水平也呈现极显著上升趋势(P<0.01)。
2.3 T-5224和KN-62刺激鳜脑细胞后味觉受体t1r1基因DNA甲基化水平的变化
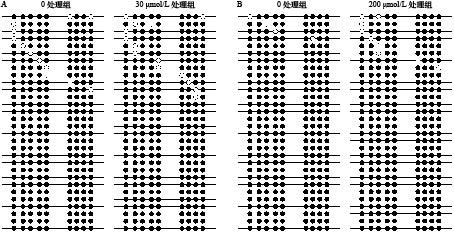
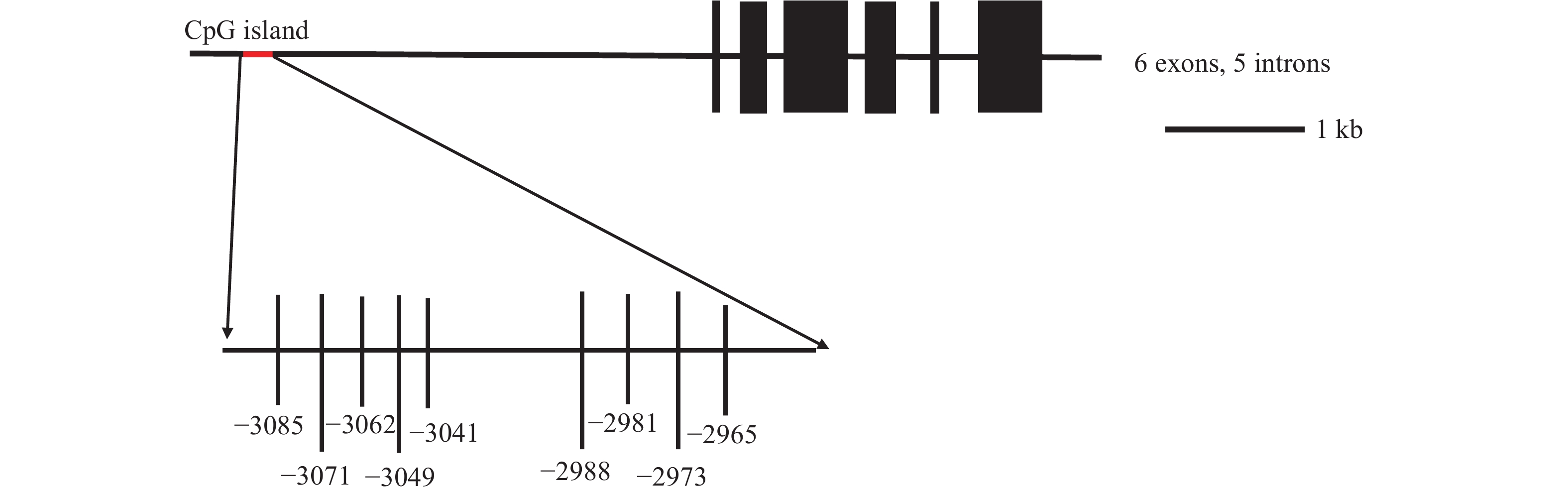
我们分析了t1r1起始密码子(指定为1)至上游–3500 bp处的CpG岛。在翘嘴鳜t1r1基因的这一区域中存在一个CpG岛, CpG 岛全长为 177 bp。从t1r1基因结构图中看到预测 CpG 岛中 9 个候选 CpG 位点的分布情况, 它们分别位于–3085、–3071、–3062、–3049、–3041、–2988、–2981、–2973 和–2965 bp (图 3)。通过前期抑制剂浓度和处理时间的最适筛选, 抑制剂T-5224 30 µmol/L和KN-62 200 µmol/L处理发现, T-5224处理组与对照组各CpG位点发生甲基化甲基化的概率很高, 但都无显著性差异, 而KN-62处理组与对照组各CpG位点发生甲基化甲基化的概率也很高, 但是也都无显著差异(表 2; 图 4)。
表 2 不同处理后t1r1的5′-侧翼区域中CpG岛中每个CpG位点的甲基化概率Table 2. Methylation status of each CpG (cytosine-guanine) site in the CpG island in the 5′-flanking regions of t1r1 (taste 1 receptor member 1) after different treatmentCpG 位点Position T-5224处理T-5224 treatment KN-62处理KN-62 treatment 0处理组(%) 200 µmol/L处理组(%) 统计(%) 显著性 0处理组(%) 200 µmol/L处理组(%) 统计(%) 显著性 –3085 96.7 90 93.3 0.612 90 90 90 1 –3071 100 96.7 98.3 1 96.7 90 93.3 0.612 –3062 96.7 90 93.3 0.612 96.7 100 98.3 1 –3049 96.7 100 98.3 1 96.7 96.7 96.7 1 –3041 100 100 100 1 93.3 93.3 93.3 1 –2988 100 96.7 98.3 1 93.3 96.7 95 1 –2981 96.7 100 98.3 1 100 96.7 98.3 1 –2973 100 100 100 1 100 93.3 96.7 0.4915 –2965 100 96.7 98.3 1 93.3 96.7 95 1 统计Total (%) 98.5 96.7 97.6 0.261 95.6 94.8 95.2 0.8412 ![]() 图 4 翘嘴鳜不同浓度抑制剂处理后t1r1基因各个CpG位点DNA甲基化水平A代表T-5224处理; B代表KN-62处理; ●代表甲基化位点, 〇代表未甲基化位点Figure 4. DNA methylation level of each CpG site of t1r1 gene treated with different concentrations of inhibitors in Siniperca chuatsiA. T-5224 processing; B. KN-62 treatment; Filled (black) circles correspond to methylated Cs, and unfilled (white) circles correspond to unmethylated Cs
图 4 翘嘴鳜不同浓度抑制剂处理后t1r1基因各个CpG位点DNA甲基化水平A代表T-5224处理; B代表KN-62处理; ●代表甲基化位点, 〇代表未甲基化位点Figure 4. DNA methylation level of each CpG site of t1r1 gene treated with different concentrations of inhibitors in Siniperca chuatsiA. T-5224 processing; B. KN-62 treatment; Filled (black) circles correspond to methylated Cs, and unfilled (white) circles correspond to unmethylated Cs3. 讨论
3.1 学习记忆因子c-fos对翘嘴鳜味觉受体t1r1基因调控表达分析
本研究结果表明抑制剂T-5224在10—30 μmol/L对抑制c-fos的转录水平效果最佳, 与此同时t1r1的转录水平也受到了抑制效果。而抑制剂KN-62的结果相反, KN-62处理后c-fos及t1r1的表达水平都显著升高。T-5224作为特异性抑制c-Fos/c-Jun的DNA结合活性而不影响包括NF-κB在内的其他转录因子的结合活性的抑制剂[28, 29], 其可以抑制c-fos基因mRNA的转录表达[30]。同时KN-62作为CaMKII的特异性激酶, 可以使得CaMKII不能发生磷酸化而处于无活性状态[31], 同时研究报道由CaMKII引起磷酸化后的CREB能引发基因表达[32], 而磷酸化后被激活的CREB参与c-fos基因的转录[33], 我们结果与此一致。有研究报道, c-fos的表达是小鼠海马区依赖性非空间记忆的形成和反复记忆所必需的[15], 而学习记忆可以影响鱼类摄食及食性的过程[34]。我们之前的研究表明, c-fos可能是影响鱼类驯食的关键基因[23]。而味觉受体t1r1就是驯食基因之一, 它不仅是鲜味感觉受体, 在动物鲜活饵料识别过程中发挥关键作用, 还在最近的研究中被发现可结合神经递质在脑中海马区表达, 与学习记忆相关[35-38]。窦亚琪等[25]通过翘嘴鳜活体研究表明c-fos的变化可能影响味觉受体t1r1的mRNA的转录水平, 从而影响鳜鱼驯化后接受人工饲料。本研究首次在翘嘴鳜脑细胞证实了抑制剂T-5224对c-fos的负调控和KN-62对c-fos的正调控, 同时表明c-fos作为一个转录因子很可能调控了翘嘴鳜味觉受体基因t1r1,从而影响其摄食偏好。
3.2 学习记忆因子c-fos对翘嘴鳜味觉受体t1r1基因DNA甲基化水平分析
DNA甲基化可能参与到精确调节基因表达使物种适应环境[38]。DNA甲基化调控机制已在多个领域中进行研究, 包括生态毒理学, 性发育和遗传育种[40-42], 然而在摄食性状与记忆形成之间联系的方面尚没有任何研究[43]。大熊猫t1r1的假基因化与大熊猫从肉食性向草食性的转化密切相关[17, 18], 进一步有研究发现, 在不同程度驯食翘嘴鳜研究中发现, 鳜脑中c-fos转录水平变化的同时, t1r1基因调控区的DNA甲基化水平也发生变化, 从而造成t1r1基因的转录水平变化。T1r1的DNA甲基化对鳜摄食死饵料鱼的记忆再巩固和稳定至关重要, 因此c-fos可能通过t1r1基因DNA甲基化参与调控t1r1转录水平[25]。本研究结果中t1r1基因在两种抑制剂的处理下其各位点及总体甲基化的概率都较高, 但是基本无差异。与窦亚琪等[25]的结果不一致, 这种差异可能是由于体外实验与体内实验的潜在差异造成的。在细胞水平上, 鳜脑细胞中c-fos可能通过未知其他途径调控t1r1的转录水平。
4. 结论
综上所述, 本研究我们比较了两种抑制剂T-5224和KN-62对翘嘴鳜脑味觉受体t1r1基因mRNA表达及DNA甲基化的影响, 结果表明c-fos可能影响味觉受体t1r1的转录水平, 但不通过t1r1基因DNA甲基化影响其转录水平。同时我们首次在鱼类上证实T-5224对c-fos的负调控及KN-62对c-fos的正调控作用。为进一步了解学习记忆促进鱼类新食性形成的作用机制, 并通过有效提高鳜鱼学习记忆能力来解决鳜鱼饲料养殖过程中难驯化及驯化不稳定问题提供了理论依据。
-
图 4 翘嘴鳜不同浓度抑制剂处理后t1r1基因各个CpG位点DNA甲基化水平
A代表T-5224处理; B代表KN-62处理; ●代表甲基化位点, 〇代表未甲基化位点
Figure 4. DNA methylation level of each CpG site of t1r1 gene treated with different concentrations of inhibitors in Siniperca chuatsi
A. T-5224 processing; B. KN-62 treatment; Filled (black) circles correspond to methylated Cs, and unfilled (white) circles correspond to unmethylated Cs
表 1 实时荧光定量 PCR 的引物序列
Table 1 Nucleotide sequences of the primers for real-time PCR
基因
Gene name引物
Primer序列
Sepuences (5′—3′)退火温度
Annealing temperature (℃)产物大小
Product size (bp)rpl13a rpl13a-F TATCCCCCCACCCTATGACA 59 100 rpl13a-R ACGCCCAAGGAGAGCGAACT c-fos c-fos-F CGATGATGTTTACCGCTTTC 60 88 c-fos-R TAGTATCCCAGATTGTCCC t1r1 t1r1-F CGAGAGTGCCAAAGGAGGAG 58 278 t1r1-R TGGAGAGCAGATGGACAGC t1r1 BSP t1r1 F AGGGTTAATATAGATATAGATAACGATAGA 52 298 BSP t1r1 R CAACTAAATAATCAATTAAAAAATACAC 表 2 不同处理后t1r1的5′-侧翼区域中CpG岛中每个CpG位点的甲基化概率
Table 2 Methylation status of each CpG (cytosine-guanine) site in the CpG island in the 5′-flanking regions of t1r1 (taste 1 receptor member 1) after different treatment
CpG 位点Position T-5224处理T-5224 treatment KN-62处理KN-62 treatment 0处理组(%) 200 µmol/L处理组(%) 统计(%) 显著性 0处理组(%) 200 µmol/L处理组(%) 统计(%) 显著性 –3085 96.7 90 93.3 0.612 90 90 90 1 –3071 100 96.7 98.3 1 96.7 90 93.3 0.612 –3062 96.7 90 93.3 0.612 96.7 100 98.3 1 –3049 96.7 100 98.3 1 96.7 96.7 96.7 1 –3041 100 100 100 1 93.3 93.3 93.3 1 –2988 100 96.7 98.3 1 93.3 96.7 95 1 –2981 96.7 100 98.3 1 100 96.7 98.3 1 –2973 100 100 100 1 100 93.3 96.7 0.4915 –2965 100 96.7 98.3 1 93.3 96.7 95 1 统计Total (%) 98.5 96.7 97.6 0.261 95.6 94.8 95.2 0.8412 -
[1] Guillette L M, Morgan K V, Hall Z J, et al. Food preference and copying behaviour in zebra finches, Taeniopygia guttata [J]. Behavioural Processes, 2014(109): 145-150. doi: 10.1016/j.beproc.2014.04.013
[2] Brown C, Laland K. Social enhancement and social inhibition of foraging behaviour in hatchery-reared Atlantic salmon [J]. Journal of Fish Biology, 2002, 61(4): 987-998. doi: 10.1111/j.1095-8649.2002.tb01857.x
[3] Brown C, Laland K N. Social learning in fishes: a review [J]. Fish and Fisheries, 2003, 4(3): 280-288. doi: 10.1046/j.1467-2979.2003.00122.x
[4] Magnhagen C, Staffan F. Social learning in young-of-the-year perch encountering a novel food type [J]. Journal of Fish Biology, 2003, 63(3): 824-829. doi: 10.1046/j.1095-8649.2003.00189.x
[5] Pike T W, Kendal J R, Rendell L E, et al. Learning by proportional observation in a species of fish [J]. Behavioral Ecology, 2010, 21(3): 570-575. doi: 10.1093/beheco/arq025
[6] Webster M M, Laland K N. Social information use and social learning in non-grouping fishes [J]. Behavioral Ecology, 2017, 28(6): 1547-1552. doi: 10.1093/beheco/arx121
[7] Webster M M, Whalen A, Laland K N. Fish pool their experience to solve problems collectively [J]. Nature Ecology & Evolution, 2017, 1(5): 135.
[8] Huber L, Range F, Voelkl B, et al. The evolution of imitation, what do the capacities of non-human animals tell us about the mechanisms of imitation [J]? Philosophical Transactions of the Royal Society of London Series B, Biological Sciences, 2009, 364(1528): 2299-2309. doi: 10.1098/rstb.2009.0060
[9] Braithwaite V A, Salvanes A G V. Environmental variability in the early rearing environment generates behaviourally flexible cod, implications for rehabilitating wild populations [J]. Proceedings Biological Sciences, 2005, 272(1568): 1107-1113.
[10] Brown G E. Learning about danger: chemical alarm cues and local risk assessment in prey fishes [J]. Fish and Fisheries, 2003, 4(3): 227-234. doi: 10.1046/j.1467-2979.2003.00132.x
[11] Hoare D J, Krause J. Social organisation, shoal structure and information transfer [J]. Fish and Fisheries, 2003, 4(3): 269-279. doi: 10.1046/j.1467-2979.2003.00130.x
[12] Ryer C H, Olla B L. Information transfer and the facilitation and inhibition of feeding in a schooling fish [J]. Environmental Biology of Fishes, 1991, 30(3): 317-323. doi: 10.1007/BF02028847
[13] Chapman B B, Ward A J W, Krause J. Schooling and learning: early social environment predicts social learning ability in the guppy, Poecilia reticulata [J]. Animal Behaviour, 2008, 76(3): 923-929. doi: 10.1016/j.anbehav.2008.03.022
[14] Sundström L F, Johnsson J I. Experience and social environment influence the ability of young brown trout to forage on live novel prey [J]. Animal Behaviour, 2001, 61(1): 249-255. doi: 10.1006/anbe.2000.1593
[15] Countryman R A, Kaban N L, Colombo P J. Hippocampal c-fos is necessary for long-term memory of a socially transmitted food preference [J]. Neurobiolog of Learning and Memory, 2005, 84(3): 175-183. doi: 10.1016/j.nlm.2005.07.005
[16] Alberini C M. Transcription factors in long-term memory and synaptic plasticity [J]. Physiological Reviews, 2009, 89(1): 121-145. doi: 10.1152/physrev.00017.2008
[17] Li R, Fan W, Tian G, et al. The sequence and de novo assembly of the giant Panda genome [J]. Nature, 2010, 463(7279): 311-317. doi: 10.1038/nature08696
[18] Zhao H, Yang J R, Xu H, et al. Pseudogenization of the umami taste receptor gene Tas1r1 in the giant Panda coincided with its dietary switch to bamboo [J]. Molecular Biology and Evolution, 2010, 27(12): 2669-2673. doi: 10.1093/molbev/msq153
[19] 刘寒. 基于基因组学的团头鲂植食性机制研究 [D]. 武汉: 华中农业大学, 2016. Liu H. Based on the genomics of Megalobrama amblycephala reveals the adaption to herbivorous diet [D]. Wuhan: Huazhong Agricultural University, 2016.
[20] Bird A. DNA methylation patterns and epigenetic memory [J]. Genes & Development, 2002, 16(1): 6-21.
[21] Plass C, Soloway P D. DNA methylation, imprinting and cancer [J]. European Journal of Human Genetics, 2002, 10(1): 6-16. doi: 10.1038/sj.ejhg.5200768
[22] 梁旭方. 翘嘴鳜人工饲料的研究 [J]. 水产科技情报, 2002, 29(2): 64-67. doi: 10.3969/j.issn.1001-1994.2002.02.001 Liang X F. The study on artificial feed of Chinese perch (Siniperca chuatsi) [J]. Fisheries Science & Technology Information, 2002, 29(2): 64-67. doi: 10.3969/j.issn.1001-1994.2002.02.001
[23] Peng J, Dou Y Q, Liang H, et al. Social learning of acquiring novel feeding habit in mandarin fish (Siniperca chuatsi) [J]. International Journal of Molecular Sciences, 2019, 20(18): 4399. doi: 10.3390/ijms20184399
[24] He S, Liang X F, Sun J, et al. Insights into food preference in hybrid F1 of Siniperca chuatsi (♀) × Siniperca scherzeri (♂) mandarin fish through transcriptome analysis [J]. BMC Genomics, 2013(14): 601. doi: 10.1186/1471-2164-14-601
[25] Dou Y, He S, Liang X F, et al. Memory function in feeding habit transformation of mandarin fish (Siniperca chuatsi) [J]. International Journal of Molecular Sciences, 2018, 19(4): 1254. doi: 10.3390/ijms19041254
[26] 石林杰, 梁旭方, 何珊, 等. 鳜脑神经元的原代培养与鉴定 [J]. 水产学报, 2020, 44(1): 43-48. Shi L J, Liang X F, He S, et al. Primary culture and identification of brain neurons in Chinese perch (Siniperca chuatsi) [J]. Journal of Fisheries of China, 2020, 44(1): 43-48.
[27] He S, Li L, Lv L Y, et al. Mandarin fish (Sinipercidae) genomes provide insights into innate predatory feeding [J]. Communications Biology, 2020(3): 361. doi: 10.1038/s42003-020-1094-y
[28] Ishida M, Ueki M, Morishita J, et al. T-5224, a selective inhibitor of c-Fos/activator protein-1, improves survival by inhibiting serum high mobility group box-1 in lethal lipopolysaccharide-induced acute kidney injury model [J]. Journal of Intensive Care, 2015(3): 49. doi: 10.1186/s40560-015-0115-2
[29] Aikawa Y, Morimoto K, Yamamoto T, et al. Treatment of arthritis with a selective inhibitor of c-Fos/activator protein-1 [J]. Nature biotechnology, 2008, 26(7): 817-823. doi: 10.1038/nbt1412
[30] Yuan L X, Liang J Q, Zhu Q C, et al. Gammacoronavirus avian infectious bronchitis virus and Alphacoronavirus porcine epidemic diarrhea virus exploit a cell-survival strategy via upregulation of cFOS to promote viral replication [J]. Journal of Virology, 2020, 95(4): e02107-e02120.
[31] 何小华, 陈建勇. 脂多糖、钙/钙调素依赖性蛋白激酶Ⅱ抑制剂对TNBS诱导BALB/C小鼠炎症性肠病的影响 [J]. 江西医药, 2013, 48(2): 117-120. doi: 10.3969/j.issn.1006-2238.2013.02.009 He X H, Chen J Y. The effects of LPS/KN62 pretreatment on colitis of BALB/C mice animal model induced by trinitrobenzene sulfonic acid [J]. Jiangxi Medical Journal, 2013, 48(2): 117-120. doi: 10.3969/j.issn.1006-2238.2013.02.009
[32] 张广辉, 王超峰. CREB依赖的突触长时程增强 [J]. 中国医学物理学杂志, 2011, 28(4): 2767-2770. doi: 10.3969/j.issn.1005-202X.2011.04.016 Zhang G H, Wang C F. CREB depended long term potentiation [J]. Chinese Journal of Medical Physics, 2011, 28(4): 2767-2770. doi: 10.3969/j.issn.1005-202X.2011.04.016
[33] Sheng M, Thompson M A, Greenberg M E. CREB: a Ca2+-regulated transcription factor phosphorylated by calmodulin-dependent kinases [J]. Science, 1991, 252(5011): 1427-1430. doi: 10.1126/science.1646483
[34] Diéguez C, Vazquez M J, Romero A, et al. Hypothalamic control of lipid metabolism: focus on leptin, ghrelin and melanocortins [J]. Neuroendocrinology, 2011, 94(1): 1-11. doi: 10.1159/000328122
[35] Ren X, Zhou L, Terwilliger R, et al. Sweet taste signaling functions as a hypothalamic glucose sensor [J]. Frontiers in Integrative Neuroscience, 2009(3): 12.
[36] Treesukosol Y, Smith K R, Spector A C. The functional role of the T1R family of receptors in sweet taste and feeding [J]. Physiology & Behavior, 2011, 105(1): 14-26.
[37] Voigt A, Bojahr J, Narukawa M, et al. Transsynaptic tracing from taste receptor cells reveals local taste receptor gene expression in gustatory Ganglia and brain [J]. The Journal of Neuroscience, 2015, 35(26): 9717-9729. doi: 10.1523/JNEUROSCI.0381-15.2015
[38] Wang Y, Leung F C C. GC content increased at CpG flanking positions of fish genes compared with sea squirt orthologs as a mechanism for reducing impact of DNA methylation [J]. PLoS One, 2008, 3(11): e3612. doi: 10.1371/journal.pone.0003612
[39] Xiao J, Song C, Liu S, et al. DNA methylation analysis of allotetraploid hybrids of red crucian carp (Carassius auratus red var.) and common carp (Cyprinus carpio L.) [J]. PLoS One, 2013, 8(2): e56409. doi: 10.1371/journal.pone.0056409
[40] Pierron F, Baillon L, Sow M, et al. Effect of low-dose cadmium exposure on DNA methylation in the endangered European eel [J]. Environmental Science & Technology, 2014, 48(1): 797-803.
[41] Wen A Y, You F, Sun P, et al. CpG methylation of dmrt1 and cyp19a promoters in relation to their sexual dimorphic expression in the Japanese flounder Paralichthys olivaceus [J]. Journal of Fish Biology, 2014, 84(1): 193-205. doi: 10.1111/jfb.12277
[42] Xing H, Wang C, Wu H, et al. Effects of atrazine and chlorpyrifos on DNA methylation in the brain and gonad of the common carp [J]. Comparative Biochemistry and Physiology Part C: Toxicology & Pharmacology, 2015, 168: 11-19.
[43] Martinowich K, Hattori D, Wu H, et al. DNA methylation-related chromatin remodeling in activity-dependent BDNF gene regulation [J]. Science, 2003, 302(5646): 890-893. doi: 10.1126/science.1090842



 下载:
下载: